An Epigenetic Mechanism Underlying Chromosome 17p Deletion-Driven Tumorigenesis
Abstract
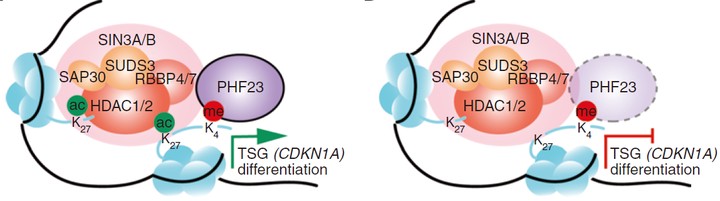
Chromosome copy-number variations are a hallmark of cancer. Among them, the prevalent chromosome 17p deletions are associated with poor prognosis and can promote tumorigenesis more than TP53 loss. Here, we use multiple functional genetic strategies and identify a new 17p tumor suppressor gene (TSG), plant homeodomain finger protein 23 (PHF23). Its deficiency impairs B-cell differentiation and promotes immature B-lymphoblastic malignancy. Mechanistically, we demonstrate that PHF23, an H3K4me3 reader, directly binds the SIN3-HDAC complex through its N-terminus and represses its deacetylation activity on H3K27ac. Thus, the PHF23-SIN3-HDAC (PSH) complex coordinates these two major active histone markers for the activation of downstream TSGs and differentiation-related genes. Furthermore, dysregulation of the PSH complex is essential for the development and maintenance of PHF23-deficient and 17p-deleted tumors. Hence, our study reveals a novel epigenetic regulatory mechanism that contributes to the pathology of 17p-deleted cancers and suggests a susceptibility in this disease.
Code and Data availability The RNA-seq and ChIP-seq data sets can be accessed under GEO accession number GSE145190. The process files and code can be found in GitHub.